Reshaping multiple variables into tidy data (wide to long)
There’s some explanation on what reshaping data in R means, why we do it, as well as the history, e.g., melt() vs gather() vs pivot_longer() in a previous post: New intuitive ways for reshaping data in R
That post shows how to reshape a single variable that had been recorded/entered across multiple different columns. But if multiple different variables are recorded over multiple different columns, then this is what you might want to do:
Example data
# from dput():
widedata = structure(list(id = c(1, 2, 3, 4, 5, 6, 7, 8, 9),
time_1 = c(1, 1, 1, 1, 1, 1, 1, 1, 1),
time_2 = c(2, 2, 2, 2, 2, 2, 2, 2, 2),
time_3 = c(3, 3, 3, 3, 3, 3, 3, 3, 3),
time_4 = c(4, 4, 4, 4, 4, 4, 4, 4, 4),
outcome_1 = c(1, 1, 1, 1, 1, 1, 1, 1, 1),
outcome_2 = c(2, 2, 2, 2, 2, 2, 2, 2, 2),
outcome_3 = c(3, 3, 3, 3, 3, 3, 3, 3, 3),
outcome_4 = c(4, 4, 4, 4, 4, 4, 4, 4, 4)),
row.names = c(NA, -9L), class = c("tbl_df", "tbl", "data.frame"))This is what it looks like:
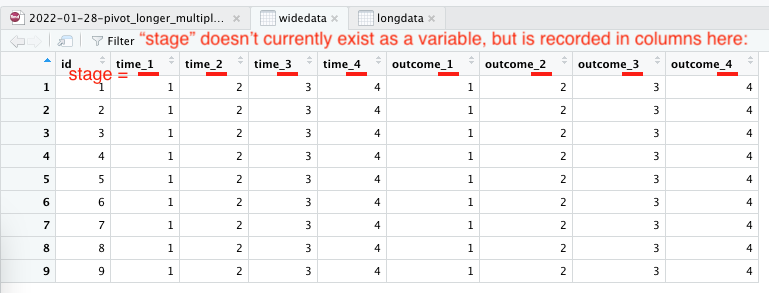
wide data in RStudio View
And this is what we want it to look like:
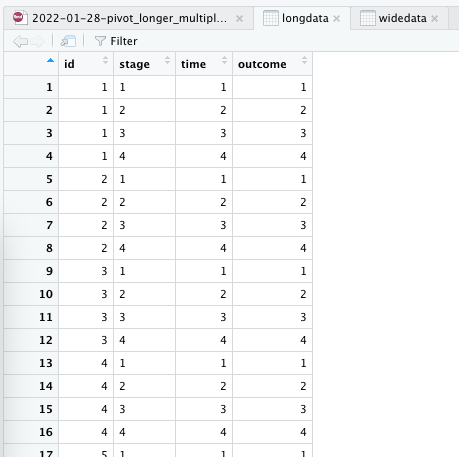
long data in RStudio View
Simple solution
library(tidyverse)
# pivot_longer puts everything that is not id into two columns (name and value)
# separate the variable names and stage numbers
# puts both variables back into two columns using pivot_wider()
longdata = widedata %>%
pivot_longer(-id) %>%
separate(name, into = c("name", "stage"), sep = "_") %>%
pivot_wider(names_from = "name", values_from = "value")I think this is a very good solution as it’s easy to understand and debug. See below for a step by step explanation of these lines.
It is, however, possible to achieve the same result just by using clever arguments inside the pivot_longer() (thank you Lisa for teaching me this):
Clever solution
# same result as above but just pivot_longer()
widedata %>%
pivot_longer(-id,
names_to = c(".value", "number"),
names_pattern = "(.+)_(.+)")# # A tibble: 36 × 4
# id number time outcome
# <dbl> <chr> <dbl> <dbl>
# 1 1 1 1 1
# 2 1 2 2 2
# 3 1 3 3 3
# 4 1 4 4 4
# 5 2 1 1 1
# 6 2 2 2 2
# 7 2 3 3 3
# 8 2 4 4 4
# 9 3 1 1 1
# 10 3 2 2 2
# # … with 26 more rowsSimple solution step by step
Combine all into two columns first
pivot_longer(-id) combines all columns except id into two columns (name and value):
widedata %>%
pivot_longer(-id)# # A tibble: 72 × 3
# id name value
# <dbl> <chr> <dbl>
# 1 1 time_1 1
# 2 1 time_2 2
# 3 1 time_3 3
# 4 1 time_4 4
# 5 1 outcome_1 1
# 6 1 outcome_2 2
# 7 1 outcome_3 3
# 8 1 outcome_4 4
# 9 2 time_1 1
# 10 2 time_2 2
# # … with 62 more rowsThe defaults name and value may be changed using these arguments:
widedata %>%
pivot_longer(-id, names_to = "my_variables", values_to = "my_values") %>% slice(1)# # A tibble: 1 × 3
# id my_variables my_values
# <dbl> <chr> <dbl>
# 1 1 time_1 1- using
%>% slice(1)for brevity
If you have multiple columns that you don’t want collected, then it’s easier to select the ones you want. In this example, it would look like this:
widedata %>%
pivot_longer(matches("time|outcome")) %>% slice(1)# # A tibble: 1 × 3
# id name value
# <dbl> <chr> <dbl>
# 1 1 time_1 1In this example dataset, matches("time|outcome") has the same effect as -id.
Search for “tidyverse select helpers” to see the various options available for selecting the variables you need (select helpers reference).
Separate column names and numbers
widedata %>%
pivot_longer(-id) %>%
separate(name, into = c("name", "stage"), sep = "_") %>% slice(1)# # A tibble: 1 × 4
# id name stage value
# <dbl> <chr> <chr> <dbl>
# 1 1 time 1 1pivot_wider() so that each variable has its own column
And the final step of this solution is pivot_wider() which takes the multiple variables that pivot_longer() combined into name and puts them into their own columns:
widedata %>%
pivot_longer(-id) %>%
separate(name, into = c("name", "stage"), sep = "_") %>%
pivot_wider(names_from = "name", values_from = "value")# # A tibble: 36 × 4
# id stage time outcome
# <dbl> <chr> <dbl> <dbl>
# 1 1 1 1 1
# 2 1 2 2 2
# 3 1 3 3 3
# 4 1 4 4 4
# 5 2 1 1 1
# 6 2 2 2 2
# 7 2 3 3 3
# 8 2 4 4 4
# 9 3 1 1 1
# 10 3 2 2 2
# # … with 26 more rowsWhat if my columns don’t have delimiters (e.g., instead of time_1, time_2,... it’s time1, time2, ...)
In that case I would use mutate() + str_extract()/str_remove():
widedata %>%
pivot_longer(-id) %>%
mutate(stage = str_extract(name, "[:digit:]")) %>%
mutate(variable = str_remove(name, "_[:digit:]")) %>%
pivot_wider(names_from = "name", values_from = "value")The first mutate extracts the number ("[:digit:]") from the column called name (which is the result of pivot_longer()), I’ve called the new variable that gets this number stage but you can call it anything. We then remove the number from name as it now lives in a column of its own.
This extraction is actually what the tidyr::extract() function is for, but it always takes me much longer to get extract() to work as compared to the easy to manage mutate() + str_extract()/str_remove() combo above.
Final words
Reshaping data is really tricky, and your spreadsheet from hell is likely much more complicated than the simple example here. It always takes me lots of trial and error to get these things to work properly. Especially if there are irregularities in the data. You may find janitor::clean_names() useful, or you may need to do more cleanup using various str_() functions from the stringr package.
Good luck!